(Very brief) Introduction
For an introduction to joint dimension reduction and clustering of continuous data, read the first part of this series.
The clustrd package contains three main functions: cluspca(), clusmca() and cluspcamix() and three auxiliary functions for cluster validation, namely tuneclus(), global_bootclus() and local_bootclus(). In this post, we will illustrate the use of clusmca(), a function that implements joint dimension reduction and clustering methods suitable for categorical data.
Cluster Correspondence Analysis on the cmc data set
Our aim is to cluster a set of observations described by categorical variables. The cmc data set refers to a survey in 1,473 married women from Indonesia who were not pregnant (or did not know they were pregnant) at the time of the survey. The data set contains nine socio-economic characteristics of the women along with their preferred method of contraception (no use, long-term methods, short-term methods). This data set is routinely used for a classification task, where the aim is to predict the contraceptive method choice of a woman, based on the nine characteristics. In the current context, however, the goal of the analysis is to identify distinct groups of Indonesian women based on their socio-economic characteristics, along with their choice of contraceptive method.
We begin with installing and loading the package and the data set:
# installing clustrd
install.packages("clustrd")
# loading clustrd
library(clustrd)
data(cmc)
summary(cmc)
W_AGE W_EDU H_EDU NCHILD
16-26:425 low :152 low : 44 0 : 97
27-39:708 2 :334 2 :178 1-4 :811
40-49:340 3 :410 3 :352 5 and above:565
high:577 high:899
W_REL W_WORK H_OCC SOL MEDEXP
non-Islam: 220 no : 369 1:436 low :129 good :1364
Islam :1253 yes:1104 2:425 2 :229 not good: 109
3:585 3 :431
4: 27 high:684
CM
no use :629
long-term :333
short-term:511First, two out of the ten variables, wife’s age in years (continuous) and number of children (count) were categorized into three groups based on quartiles.
# Data preprocessing: values of wife's age and number of children were categorized
# into three groups based on quartiles
cmc$W_AGE = ordered(cut(cmc$W_AGE, c(16,26,39,49), include.lowest = TRUE))
levels(cmc$W_AGE) = c("16-26","27-39","40-49")
cmc$NCHILD = ordered(cut(cmc$NCHILD, c(0,1,4,17), right = FALSE))
levels(cmc$NCHILD) = c("0","1-4","5 and above")The clusmca() function has three arguments that must be provided: a data.frame (data) with categorical variables, the number of clusters (nclus) and the number of dimensions (ndim). Optional arguments include method, alphak, nstart, smartStart, gamma, binary and seed, and are described in help(clusmca). Our method of choice in this example is Cluster Correspondence Analysis (van de Velden, Iodice D’Enza, & Palumbo 2017), which combines Correspondence Analysis for dimension reduction with K-means for clustering. The method argument is set to “clusCA”.
For instance, the solution with three clusters and two dimensions with 100 random starts is given by:
out.clusmca = clusmca(cmc, nclus = 3, ndim = 2, method = "clusCA", nstart = 100, seed = 1234)
out.clusmcaChoosing the number of clusters and dimensions
OK, how many clusters and how many dimensions should we select? The decision on the appropriate number of clusters and dimensions can be based on the value of the overall average Silhouette width (ASW) or the Calinski-Harabasz (CH) index of the final clusters obtained via clusmca(). The calculation of the ASW and CH can be based on a distance matrix either between the original observations (using Gower’s dissimilarity) or between their scores in the low-dimensional space (using the Euclidean distance). This procedure is implemented in the function tuneclus().
bestclusCA = tuneclus(cmc, nclusrange = 3:10, ndimrange = 2:9, method = "clusCA", criterion = "asw", nstart = 100, seed = 1234)Notice that the number of clusters varies from 3 to 10 and the number of dimensions from 2 to 9. Distances are computed between observations on the original data matrix (argument dst = “full”). The method argument is set to “clusCA” with 100 random starts (nstart = 100) and the criterion argument to “asw”. See help(tuneclus) for more Details.
Warning: the tuneclus() function will take really long time to execute; use nstart = 5 to speed up computation (the results may be slightly different, however).
The best solution was obtained for 7 clusters of sizes
392 (26.6%), 373 (25.3%), 287 (19.5%), 190 (12.9%),
113 (7.7%), 91 (6.2%), 27 (1.8%) in 6 dimensions,
for a cluster quality criterion value of 0.194.
Cluster quality criterion values across the specified
range of clusters (rows) and dimensions (columns):
2 3 4 5 6 7 8 9
3 0.177
4 0.111 0.168
5 0.099 0.113 0.12
6 0.065 0.102 0.115 0.13
7 0.052 0.072 0.098 0.097 0.194
8 0.044 0.059 0.07 0.119 0.129 0.147
9 0.045 0.057 0.057 0.111 0.114 0.108 0.163
10 0.044 0.074 0.091 0.088 0.099 0.113 0.134 0.144
Cluster centroids:
Dim.1 Dim.2 Dim.3 Dim.4 Dim.5 Dim.6
Cluster 1 0.9517 0.2099 -0.0273 0.1150 0.7146 0.1424
Cluster 2 -0.5170 0.0667 -0.4497 -0.0045 -0.0383 -0.7735
Cluster 3 -0.4522 -0.9552 -0.4419 0.4141 -0.2591 0.5546
Cluster 4 1.2383 0.5261 0.2256 -0.2934 -1.1737 0.0916
Cluster 5 -1.9431 1.4499 0.0983 -0.7524 0.1553 0.7812
Cluster 6 -0.3352 -1.3685 1.8556 -1.2789 0.2356 -0.2544
Cluster 7 -1.3194 1.0271 3.0539 3.5157 -0.2768 -0.3340
Within cluster sum of squares by cluster:
[1] 292.0676 264.6142 217.1584 200.6005 189.4055 177.0946 106.8898
(between_SS / total_SS = 74.7 %)
Objective criterion value: 966.1789
Available output:
[1] "clusobjbest" "nclusbest" "ndimbest" "critbest"
[5] "critgrid"Given that the best solution is obtained for 7 clusters and 6 dimensions, we can subsequently visualize observations, cluster centers and variable categories on the first two dimensions, similar to Multiple Correspondence Analysis.
# plotting the Cluster CA solution
plot(bestclusCA, cludesc = TRUE)This creates a scatterplot of observations, cluster centers and variable categories as shown below. Note that the argument cludesc = TRUE additionally creates a series of barplots showing the standardized residuals per variable category for each cluster to facilitate cluster interpretation (see below).
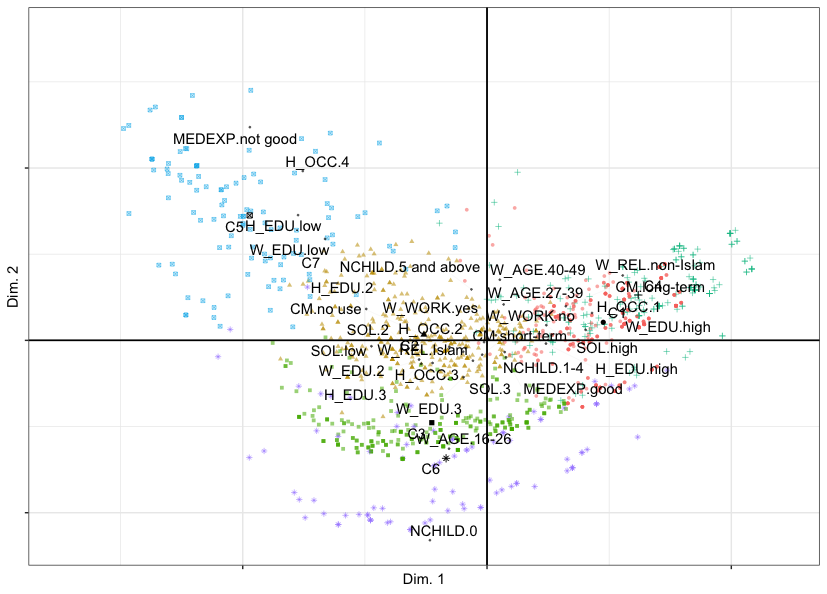
In the following figure, we can notice that Dimension 1 opposes women with low education on the left (Clusters 5 and 7) to women with high education on the right (Clusters 4 and 1) - this could be named the education dimension. Dimension 2 opposes younger women with no children (bottom, Clusters 6 and 3) to older women with 5 children or above (middle, Cluster 2) - this is the age-children dimension.
Cluster Correspondence Analysis map of observations, cluster centers and variable categories of the cmc data set with respect to components 1 (horizontal) and 2 (vertical). Cluster centers are labelled C1 through C7.
The same results could have been simply obtained via clusmca(), as follows:
out.clusmca = clusmca(cmc, 7, 6, method = "clusCA", nstart = 100, seed = 1234)
plot(out.clusmca, cludesc = TRUE)Since there are six dimensions, we can also visualize dimensions 1 and 3, 2 and 3 and so on. This can further reveal the clusters that oppose each other. For instance, let’s plot dimensions 5 and 6.
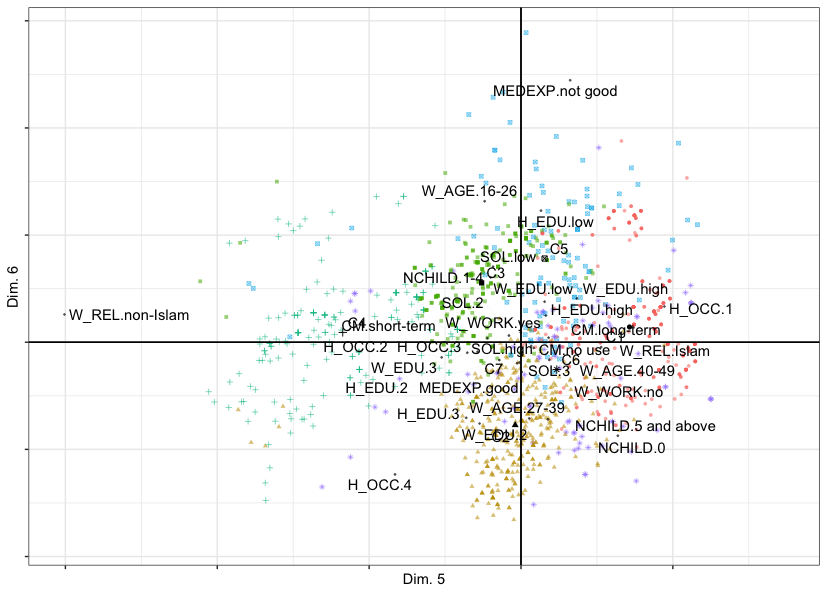
plot(out.clusmca, dims = c(5, 6))The figure below shows that Dimension 5 opposes women with non-Islamic religion on the left (Cluster 4) to women with Islamic religion on the right (Cluster 1) - this is the dimension of religion. Dimension 6 opposes women with not good media exposure (top, Cluster 5) to the rest of women (middle and bottom) - this is the dimension of media exposure.
Cluster Correspondence Analysis map of observations, cluster centers and variable categories of the cmc data set with respect to components 5 (horizontal) and 6 (vertical). Cluster centers are labelled C1 through C7.
Assessing Cluster Stability
Let’s assess how stable each cluster in the solution is. In the first part of this post series, you can find more details on Cluster stability and how it is assessed in clustrd. See also help(local_bootclus).
bootres = local_bootclus(cmc, nclus = 7, ndim = 6, method = "clusCA", nboot = 100, nstart = 50, seed = 1234)
summary(bootres$Jaccard)
1 2 3
Min. :0.4713 Min. :0.2994 Min. :0.2274
1st Qu.:0.6840 1st Qu.:0.5704 1st Qu.:0.5460
Median :0.7820 Median :0.6479 Median :0.6086
Mean :0.7660 Mean :0.6464 Mean :0.6422
3rd Qu.:0.8561 3rd Qu.:0.7874 3rd Qu.:0.8510
Max. :0.9475 Max. :0.8932 Max. :0.9193
4 5 6
Min. :0.1858 Min. :0.3337 Min. :0.4195
1st Qu.:0.5745 1st Qu.:0.5614 1st Qu.:0.8264
Median :0.6057 Median :0.6575 Median :0.8544
Mean :0.6980 Mean :0.6381 Mean :0.7918
3rd Qu.:0.9424 3rd Qu.:0.7380 3rd Qu.:0.8699
Max. :0.9818 Max. :0.8701 Max. :0.9151
7
Min. :0.1375
1st Qu.:0.3738
Median :0.4853
Mean :0.4792
3rd Qu.:0.6437
Max. :0.8864boxplot(bootres$Jaccard, xlab = "cluster number", ylab = "Jaccard similarity (cluster stability)")Local stability of a Cluster Correspondence Analysis solution with 7 clusters in 6 dimensions.
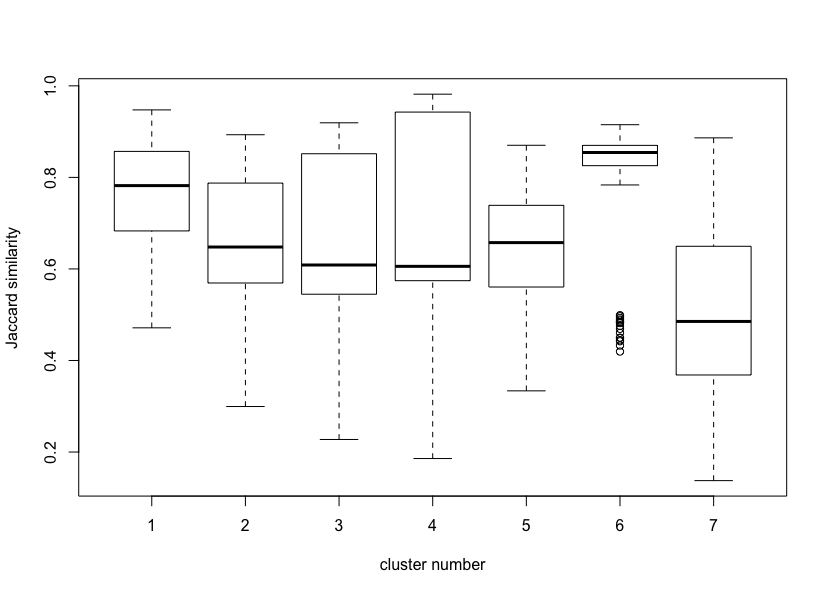
The results indicate that clusters 1 and 6 are qute stable (mean Jaccard similarities greater than 0.75), clusters 2, 3, 4 and 5 are not that stable but are indicating patterns in the data (mean Jaccard similarities between 0.6 and 0.75), and cluster 7 is not stable (below 0.6). A visual inspection of the Jaccard similarity distributions via boxplots is shown above.
Cluster Interpretation
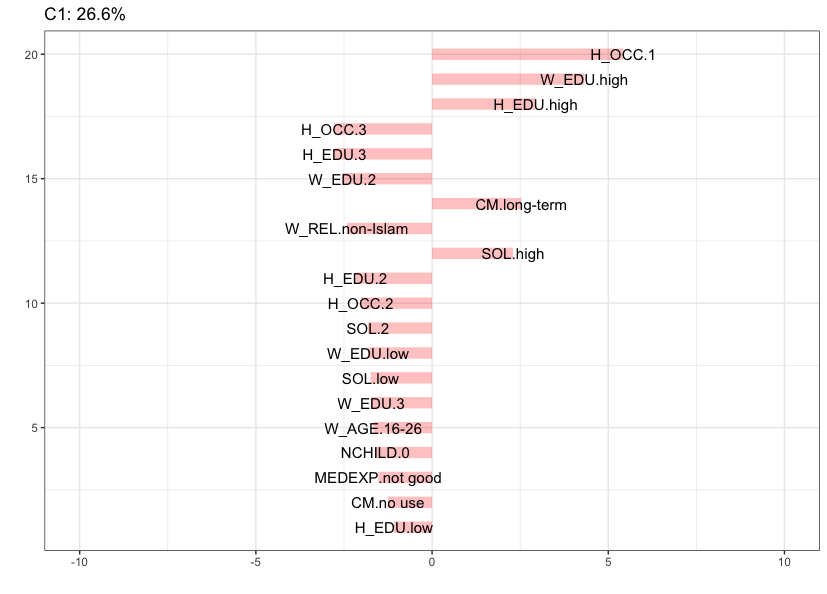
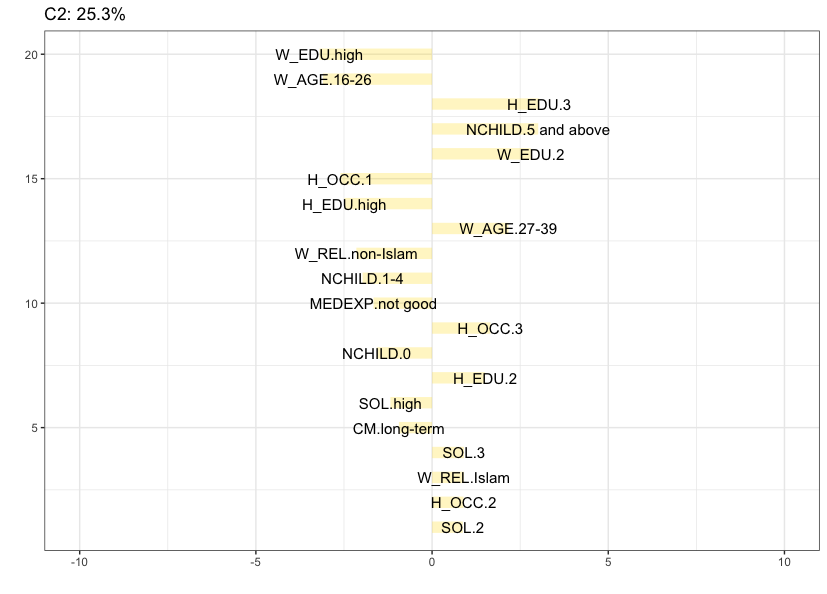
To help with the interpretation of clusters it is useful to identify attributes that deviate the most from the independence condition. The seven barplots below, obtained via the argument cludesc = TRUE, show for each cluster the twenty categories with the highest standardized residuals (positive or negative). Let’s inspect the content of each cluster.
The figure below shows that Cluster 1 (26.6%), which is the largest cluster in the data, is mainly characterized by women associated with a high education, having a husband with high education, a high standard of living, husband’s occupation is 1 and reported to be using long-term contraceptive methods. Note that the data do not contain labels for husband’s occupation. Recall that Cluster 1 is a quite stable cluster in the data.
Cluster 2 (25.3%) is also quite large and is mainly characterized by women with 5 children or more, that belong to the 27-39 age category, having low to moderate education (both wife and husband), a low to moderate standard of living, an Islamic religion and husband’s occupation is 2 or 3.
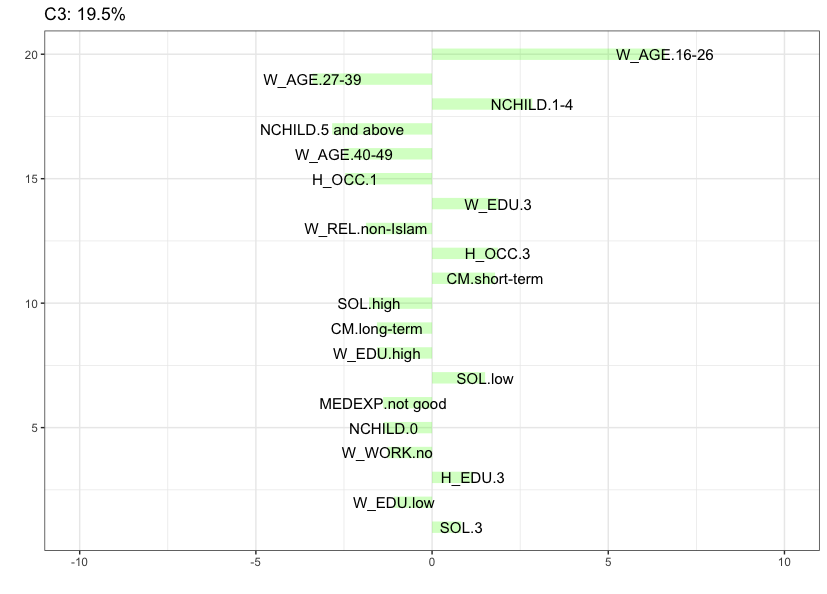
Cluster 3 (19.5%) contains women that belong to the lowest age category, having between 1 and 4 children, a moderate to high education (both wife and husband), a low to moderate standard of living, husband’s occupation is 3 and reported to be using short-term contraceptive methods.
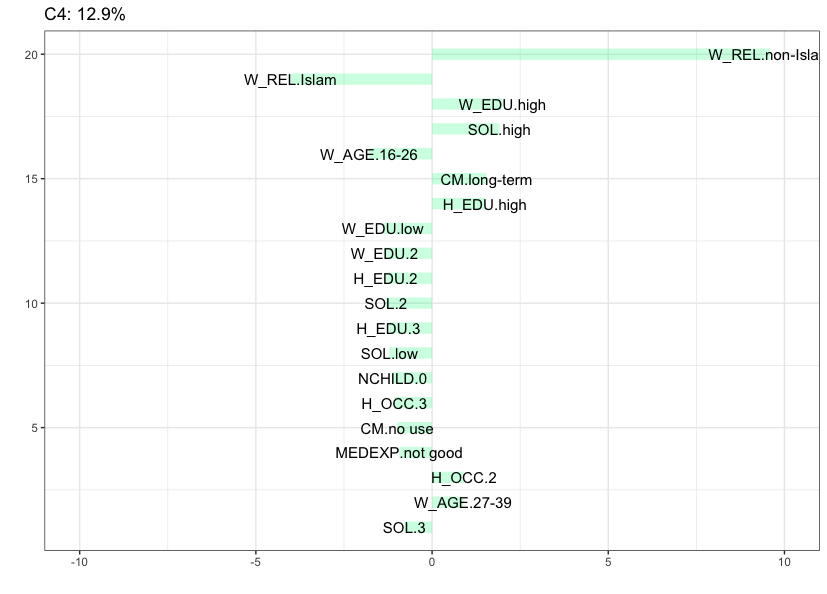
Cluster 4 (12.9%) contains women with non-Islamic religion, that belong to the 27-39 age category, have a husband with high education, a high standard of living, a good media exposure and reported to be using long-term contraceptive methods.
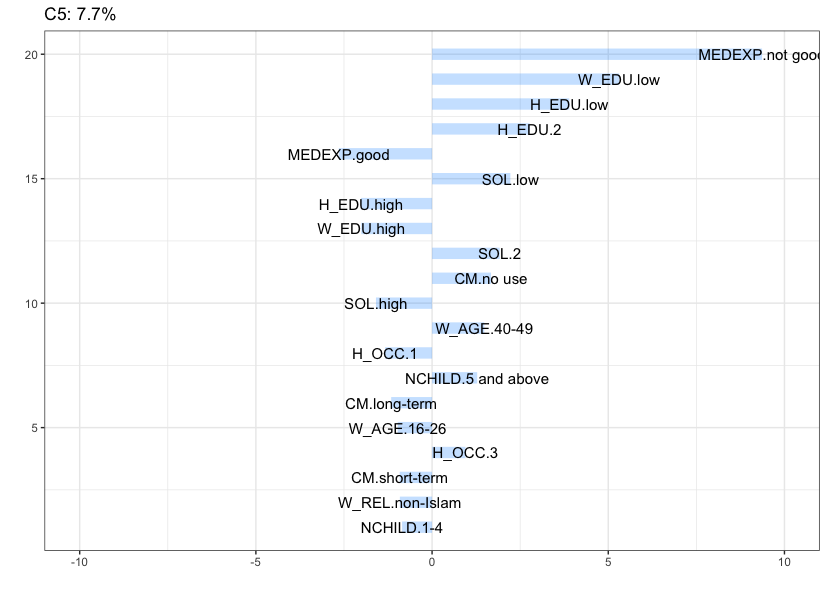
Cluster 5 (7.7%) is characterized by women with a low education level (also their husband), that belong to the highest age category (40-49), have a low standard of living, 5 children or above, and reported to be using no contraception at all.
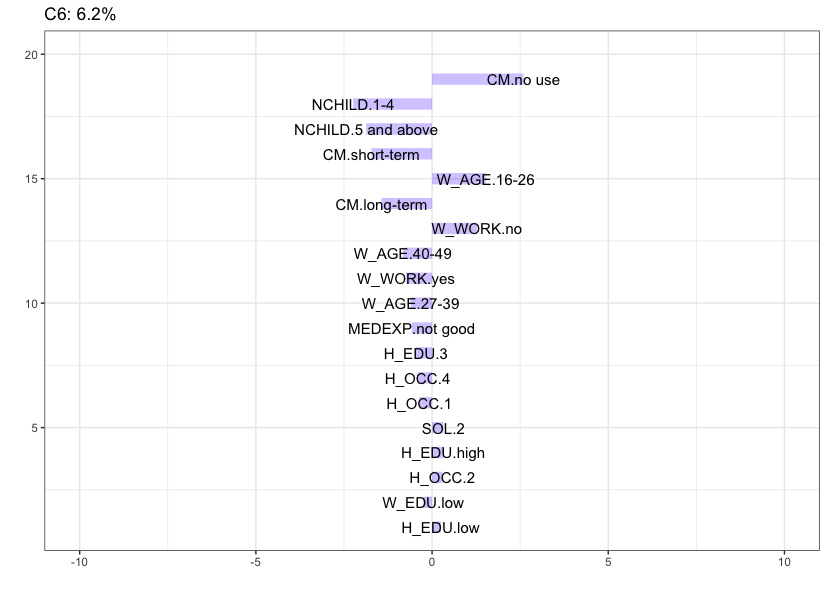
Cluster 6 (6.2%), which is the most stable cluster in the data, is characterized by women that belong to the lowest age category (16-26), do not work, have no children and reported to be using no contraception at all.
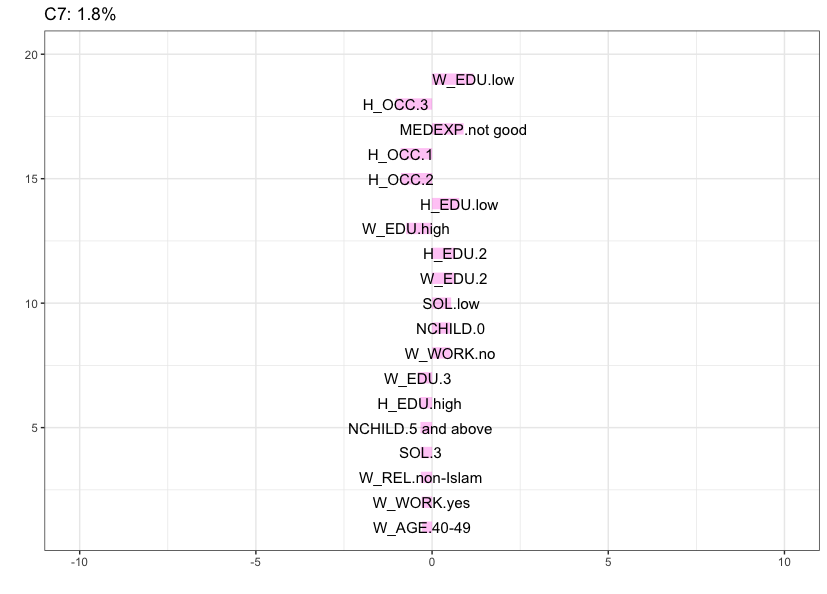
Cluster 7 (1.8%) is the smallest cluster, characterized by women in the lowest age category (16-26) that do not work and husband’s occupation is 4 (this cluster contains all the women with a husband’s occupation in this category). Recall, however, that this is not a stable cluster.
Epilogue
For a thorough treatment of joint dimension reduction and clustering methods and additional applications you can read our JSS and WIREs papers. Note, however, that cluster stability is not discussed in any of these papers, since this functionality was only added to the package later on. The clustrd package is still evolving; ongoing refinements, new features and fixing bugs are being enhanced regularly. Any feedback will be much appreciated.
In other posts, I demonstrate the application of joint dimension reduction and clustering to continuous and mixed-type data sets.
References
Arabie, P., & Hubert, L. (1994). Cluster Analysis in Marketing Research. In R. P. Bagozzi. (Ed.), Advanced Methods of Marketing Research (pp. 160-189). Cambridge: Blackwell.
De Soete, G., & Carroll, J. D. (1994). K-means clustering in a low-dimensional euclidean space. In E. Diday, Y. Lechevallier, M. Schader, P. Bertrand, & B. Burtschy (Eds.), New Approaches in Classification and Data Analysis (pp. 212-219). Springer-Verlag, Berlin.
Dolnicar, S., & Grün, B. (2008). Challenging “factor–cluster segmentation”. Journal of Travel Research, 47(1), 63-71.
Dolnicar, S., & Leisch, F. (2010). Evaluation of structure and reproducibility of cluster solutions using the bootstrap. Marketing Letters, 21(1), 83-101.
Dolnicar, S., & Leisch, F. (2017). Using segment level stability to select target segments in data-driven market segmentation studies. Marketing Letters, 28(3), 423-436.
Hennig, C. (2007). Cluster-wise assessment of cluster stability. Computational Statistics and Data Analysis, 52, 258-271.
Hwang, H., Dillon, W. R., & Takane, Y. (2006). An Extension of Multiple Correspondence Analysis for Identifying Heterogenous Subgroups of Respondents. Psychometrika, 71, 161-171.
Iodice D’Enza, A., & Palumbo, F. (2013). Iterative factor clustering of binary data. Computational Statistics, 28(2), 789-807.
Markos, A., Iodice D’Enza, A., & van de Velden, M. (2018). Beyond Tandem Analysis: Joint Dimension Reduction and Clustering in R. Journal of Statistical Software (accepted for publication).
Milligan, G. W. (1996). Clustering validation: results and implications for applied analyses. In P. Arabie, L. J. Hubert (Eds.), Clustering and classification (pp. 341-375). River Edge: World Scientific Publications.
Pfitzner, D., Leibbrandt, R., & Powers, D. (2009). Characterization and evaluation of similarity measures for pairs of clusterings. Knowledge and Information Systems, 19(3), 361-394.
van de Velden, M., Iodice D’Enza, A., & Palumbo, F. (2017). Cluster Correspondence Analysis. Psychometrika, 82(1), 158-185.
van de Velden, M., Iodice D’Enza, A., & Markos, A. (2019). Distance-based clustering of mixed data. WIREs Computational Statistics, 11(3), e1456. DOI:10.1002/wics.1456.
Vichi, M., Vicari, D., & Kiers, H.A.L. (2019). Clustering and dimension reduction for mixed variables. Behaviormetrika doi:10.1007/s41237-018-0068-6.
Vichi, M., & H. A. (2001). Factorial k-means analysis for two-way data. Computational Statistics & Data Analysis, 37(1), 49-64.
